Abstract
Fungi play a critical role in silage and rumen fluid, both of which share similar ecosystems. This study is designed to supplement sweet sorghum ensiling with the rumen fluid from beef heifers and evaluate the effect on fungal diversity dynamics and silage quality by high-throughput sequencing and chemical methods. Simultaneously, this study investigated the correlation between fungal communities and chemical compositions of silages. The results suggested that the addition of rumen fluid significantly affected the richness and diversity of fungi in silage. Alpha diversity of total fungi and the relative abundance of Pichia increased, while Schizophyllum and Penicillium decreased, following the supplementation of rumen fluid during 60 days ensiling. In addition, the silage quality was positively correlated with Pichia, but negatively correlated with Hannaella and Vishniacozyma. The findings reveal that the rumen fluid can improve the ensiling characteristics by effectively reducing the reproduction of adverse fungi. However, further research is required to verify the connection of specific pure fungal isolates with the fermentation performance of silage.
Download PDF
Full Article
Effects of Rumen Fluid as Bioaugmentation Additive on Fungal Diversity Dynamics during Sweet Sorghum Ensiling
Hui Tian,a Ruifeng Shi,a Huiyuan Wei,a Nana Lu,a Wenli Sun,a Haiwei Ren,a,* Yi Zheng,b Jingping Li,c and Xiaoli Wang d
Fungi play a critical role in silage and rumen fluid, both of which share similar ecosystems. This study is designed to supplement sweet sorghum ensiling with the rumen fluid from beef heifers and evaluate the effect on fungal diversity dynamics and silage quality by high-throughput sequencing and chemical methods. Simultaneously, this study investigated the correlation between fungal communities and chemical compositions of silages. The results suggested that the addition of rumen fluid significantly affected the richness and diversity of fungi in silage. Alpha diversity of total fungi and the relative abundance of Pichia increased, while Schizophyllum and Penicillium decreased, following the supplementation of rumen fluid during 60 days ensiling. In addition, the silage quality was positively correlated with Pichia, but negatively correlated with Hannaella and Vishniacozyma. The findings reveal that the rumen fluid can improve the ensiling characteristics by effectively reducing the reproduction of adverse fungi. However, further research is required to verify the connection of specific pure fungal isolates with the fermentation performance of silage.
DOI: 10.15376/biores.17.2.2977-2996
Keywords: Sweet sorghum; Silage; Rumen fluid; Fungus; Diversity; Correlation
Contact information: a: School of Life Sciences and Engineering, Lanzhou University of Technology/Key Laboratory of Complementary Energy System of Biomass and Solar Energy, Lanzhou, Gansu Province 730050, P. R. China; b: Department of Grain Science and Industry, Kansas State University, Manhattan, KS 66506, USA; c: Lanzhou University of Technology, Western China Research Center of Energy & Environment, Lanzhou, P. R. China; and d: Lanzhou Institute of Husbandry and Pharmaceutical Sciences, Chinese Academy of Agricultural Sciences, Lanzhou 730050, P. R. China.
* Corresponding author: rhw52571119@lut.edu.cn
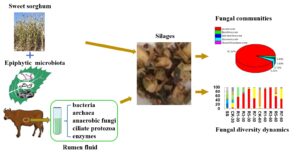
GRAPHICAL ABSTRACT

ABBREVIATIONS
AA: Acetic acid; DM: Dry matter;
ADF: Acid detergent fiber; HC: Hemicellulose;
ADL: Acid detergent lignin; LA: Lactic acid;
AN: Ammonia nitrogen; NDF: Neutral detergent fiber;
BA: Butyric acid; OTU: Operational taxonomic unit;
CL: Cellulose; WSC: Water-soluble carbohydrates
CP: Crude protein;
INTRODUCTION
As a common storage method, ensiling is effective in preserving wet forage by producing organic acids through anaerobic fermentation. After fresh raw materials are compressed and sealed, an internal anoxic environment can be formed. A long preservation time of raw materials can be realized with almost no spoilage and few losses of organic components during ensiling (Liu et al. 2018).
The silage process is initiated by natural microbiota of the forage plant, contaminants, and/or inoculants (Ávila and Carvalho 2020). Among the most abundant microbial species, lactic acid bacteria (LAB) represent the desirable bacteria, while moulds and yeasts are usually considered undesirable eukaryotes during ensiling (Ávila and Carvalho 2020). As the facultatively anaerobic bacteria, LAB multiply well and perform fermentation under the anaerobic ensiling conditions (Doyle et al. 2019; Wang et al. 2022). Generally, growth of the yeasts, molds, and some aerobic Bacillus, would lead to silage decay upon exposure to air, along with the increase of temperature and pH (Zhang et al. 2019). In essence, the success of ensiling depends on the growth of LAB and their production of lactic acid (LA) (Kung et al. 2018; Ávila and Carvalho 2020). With the lactic acid production and the decrease in pH level, growth of spoilage microorganisms containing aerobic bacteria, yeasts, and filamentous fungi can be limited (Ambye-Jensen et al. 2013). Farmers mostly rely on the natural LAB microbiota of crops to complete the ensiling process (del Palacio et al. 2016). However, the initial number of epiphytic LAB was observed to be not enough to reproduce and reduce pH value quickly to suppress undesirable microbes during ensiling (Vu et al. 2019). Thus, good field management and silage processing conditions were needed to control unfavorable microbes (Kharazian et al. 2017). In contrast, the introduction of various additives into silages could practicably ensure ensiling quality, such as salt, sugar, LAB, enzymes, and other exogenous biological agents (Vu et al. 2019; Guo et al. 2020; Ren et al. 2021). These additives could effectively improve the fermentation characteristics of silage, promote the degradation, conversion, and utilization efficiency of lignocellulose components, and inhibit the silage fungus growth (Guo et al. 2020). Nkosi et al. (2012) found that the addition of Lactobacillus buchneri (LB), Lactobacillus plantarum (LP), and LB+enzymes preparation (compound of cellulase, xylanase, and beta-glucosidase) in sweet sorghum silage could significantly increase the lactic acid concentration and reduce pH and the contents of butyric acid (BA), ammonia nitrogen (AN), and fiber (Nkosi et al. 2012). Li et al. (2019) found that ferulic acid esterase-producing LAB and cellulase pretreatments could improve the quality of corn straw silage and promote the cellulosic conversion efficiency (Li et al. 2019). Wang et al. (2022) believed that the simultaneous addition of Lactobacillus plantarum and cellulase could effectively improve the quality of mixed silage of whole-plant corn and peanut vines as well as the degradation rate of lignocellulose (Wang et al. 2022).
Recently, researchers tried to introduce isolates resourced from rumen fluid or rumen fluid itself into lignocellulosic biomass conversion, which is based on the fact that rumen fluid contains various microbes (bacteria, archaea, anaerobic rumen fungi, and ciliate protozoa) and metabolic enzymes (cellulase, hemicellulase, protease, etc.), and exhibited powerful lignocellulolytic capacity (Mackie et al. 2000; Sirohi et al. 2012; Yue et al. 2013). When treated with the rumen fluid, the lignocellulose component content of rice straw decreased significantly, with an 83% increase in methane production and a 40% reduction in the cellulose (CL) degradation time (Zhang et al. 2016). Inoculation of a rumen fungus Piromyces sp. CN6 CGMCC 14449 into corn silage caused significant reductions in pH value and contents of acetic acid (AA), acid detergent fiber (ADF), and neutral detergent fiber (NDF), while the contents of LA, crude protein (CP), and fermentable sugar increased (Wang et al. 2019).
The microorganisms of rumen fluid are complex and large-scale; only 10 to 20% of the internal microbial population have been identified through traditional culture methods, while an increasing number of microorganisms are being identified by modern high-throughput sequencing techniques (Yue et al. 2013). When using the vigorous rumen fluid during the ensiling process, fungi among the communities also play a significant role, with their advantages in penetrating and disrupting plant tissues, and then facilitating bacterial access to plant biomass (Krause et al. 2013). In contrast, some species of molds can lead to spoilage of silage or production of mycotoxins, resulting in a reduction in the silage value (del Palacio et al. 2016; Panasiuk et al. 2019).
However, information on the correlation between fungal populations and fermentation characteristics of silage is currently not enough. Thus, it is critical to perform a qualitative and quantitative assessment of the fungi communities during ensiling and to investigate the role of rumen fluid additives in the improvement of silage quality.
In the authors’ previous paper (Ren et al. 2021), a novel ensiling strategy of sweet sorghum with rumen fluid bioaugmentation was studied, focusing on the relationship between bacterial communities and ensiling performance. The objective of this study was to determine effects of rumen fluid additives on the fungal diversity of sweet sorghum silage and its correlation with fermentation characteristics and chemical components, hypothetically to efficiently improve the ensiling performance and enzymatic digestibility of silages.
EXPERIMENTAL
Materials
Sweet sorghum and rumen fluid used in this study have been described in the current authors’ previous research (Ren et al. 2021). The whole plant of sweet sorghum was harvested in the experimental plot (N 37° 93′ latitude, E 102° 35′ longitude, altitude 2200 to 2400 m a.s.L., Wuwei, China) of the Institute of Modern Physics, Chinese Academy of Sciences, and chopped into 2- to 3-cm-long particles for ensiling. Approximately 10 L of rumen fluid was obtained from fattened beef heifers, which were offered hay diets for 5 to 7 days prior to being slaughtered in the Dingle Ecological Animal Husbandry Co., Ltd. (Wuwei, China). The rumen fluid was pretreated by filtration through five-layer sterile gauze to remove suspended solid particles under aseptic conditions. The filtrate was collected and mixed thoroughly, followed by being frozen at -20 °C until used for ensiling.
Methods
Silage preparation with rumen fluid additive
Silage preparation was completed according to protocols reported in the authors’ previous study (Ren et al. 2021). Thirty pieces of 1.5 kg chopped sweet sorghum were accurately weighed and evenly sprayed with different dosages of rumen fluid [0 g (control, CK), 1 g (R1), 3 g (R3), 5 g (R5), and 7 g (R7) rumen fluid/100 g wet raw sweet sorghum]. In detail, four experimental groups of rumen fluid silage were named R1, R3, R5, and R7. The blank control group (CK) was sprayed with equal volume of distilled water. Three replicates for each treatment were packed and sealed in 30 pre-weighed plastic buckets (diameter × height = 20 × 32 cm), and then incubated at 25 ± 1 °C for 30 and 60 days ensiling.
Fungal flora analysis during ensiling fermentation
The total DNA from silage samples was extracted using the PowerSoil DNA Isolation kit (MO BIO Laboratories, Carlsbad, CA, USA) according to the method previously reported (Liu et al. 2018). Ratios of 260 nm/230 nm and 260 nm/280 nm were used to evaluate quantity and quality of the extracted DNA, and then the DNA was stored at -80 °C for sequencing analysis (Huang et al. 2021). Internal transcribed spacer (ITS) primer set ITS1F (5′-CTTGGTCATTTAGAGGAAGTAA-3′) and ITS2 (5′-GCTGCGT-TCTTCATCGATGC-3′) targeting the ITS1 region was applied to analyze the fungal communities (Li et al. 2020). The polymerase chain reaction (PCR) amplification of ITS1 region was performed as the procedure previously reported (Romero et al. 2017). The reaction programme was as follows: 95 °C for 3 min, followed by 35 cycles of 95 °C for 30 s, 55 °C for 45 s, and 72 °C for 60 s, and a final extension of 72 °C for 10 min. The PCR products were then purified by VAHTS® DNA Clean Beads (Vazyme Biotech Co., Ltd., Nanjing, China), quantified by Quant-iT™ dsDNA HS Reagent (Thermo Fisher Scientific, Waltham, MA, USA), pooled together, and finally analyzed via high-throughput sequencing technology by the Biomarker Technologies Corporation (Beijing, China) using the Illumina Hiseq 2500 platform (2 × 250 paired ends) (Illumina Inc., San Diego, CA, USA) (Huang et al. 2021).
Bioinformatic analysis of sequencing data
The final effective tags were obtained after removal of low-quality reads and chimeric sequences. The sequencing results were analyzed using Uparse software (Edgar, R.C., Version 7.0.1001, Tiburon, CA, USA) (Edgar 2013) and were compared with genes in GenBank of NCBI (National Center for Biotechnology Information, Bethesda, MD, USA). The similarity level of 97% was selected for the operational taxonomic unit (OTU) analysis. Alpha diversity indices (Shannon, Chao1, and ACE (abundance-based coverage estimator)) of samples were calculated by the QIIME software (Center for Microbiome Innovation, University of California San Diego, La Jolla, CA, USA). Fungal flora composition analysis at phylum level and genus level was performed by selecting a relative abundance higher than 0.1%. The correlation between fungal communities and chemical indices were carried out on the Biomarker (BMK) Cloud Platform (www.biocloud.net).
Correlation analysis
Chemical and fermentative characteristics during ensiling were determined using methods described in the authors’ previous report (Ren et al. 2021).
The correlations between chemical components, fermentation quality, and fungal community in the sample were analyzed using the BMK Cloud platform in the form of correlation heat map.
Data analysis
All data were statistically analyzed using one-way analysis of variance (ANOVA) with 3 replicates by IBM SPSS 20.0 (IBM Corp., Armonk, NY, USA). Statistical significance declared at P ≤ 0.05. Graphics were drawn using Origin 9.0 (OriginLab, Northampton, MA, USA).
RESULTS AND DISCUSSION
A better understanding of the fungi flora involved in the ensiling process will help to develop strategies to improve the fermentation characteristics of sweet sorghum silage. In the present study, the authors compared the fungi diversity and abundance of sweet sorghum without rumen fluid addition and with different supplementations of rumen fluid and assessed the fungi diversity dynamics from 30 to 60 days of fermentation. Moreover, correlation between fungal composition and silage quality was also analyzed.
Effect of Rumen Fluid Additive on Fungal Diversity During Ensiling
Ensiling, as an effective method for the conservation of forage and animal feedstuffs, is achieved by creating the anaerobic environment to promote lactic fermentation by LAB. In the anaerobic and acidic niche (pH 3.5 to 4.2), growth of undesirable microorganisms would be limited, which is key to animal feeding, for the microbial population could exert a direct influence on the quality and storage stability of ensiled feeds, as well as the health of animals (Drouin and Ferrero 2020; Yu et al. 2020; Ren et al. 2021).
Rumen, one of the most complex gastro-intestinal systems in ruminating animals, is an oxygen-free environment with a pH from 5.8 to 6.4 and a temperature between 37 and 42 °C, which is also well-suited for growth of anaerobic microbes. Rumen fluid inside this system is inhabited by hundreds of billions of microorganisms composed of bacteria, fungi, and protozoa (Kameshwar et al. 2019). Based on this information, inoculation of rumen fluid from healthy ruminating animals into silage in principle would not cause contamination of ensiling. Indeed, rumen fluid has proven an effective bioaugmented additive for sweet sorghum ensiling, because of its powerful lignocellulose hydrolytic and acidogenic activities (Ren et al. 2021).
Alpha diversity analysis
The alpha-diversity has been extensively used to analyze the rumen microbial communities from many kinds of ruminants, including cows, yaks, goats, and sheep (Fan et al. 2020; Langda et al. 2020; Wang et al. 2021). Based on the OTU abundance data from ITS sequencing, the Chao1 and ACE indices have been often used to unravel the total species richness in samples, while the Shannon and Simpson indices potentially provide insights on the richness as well as the evenness (Huang et al. 2021).
The diversity of fungi in seven silage samples (rumen fluid, raw sweet sorghum, CK, R1, R3, R5, and R7) were revealed by high-throughput sequencing. As a result, 873,589 reads in total of ITS1 sequences were obtained from raw MiSeq data, and then sequences were classified into 1,340 OTUs based on the 97% threshold. The alpha diversity of the fungal community based on OTUs level is shown in Table 1, consisting of the OTU, Chao1, ACE, and the Shannon and Simpson indices of different samples.
Most of the fungal communities were identified based on the coverage values (> 99%). The alpha-diversity parameters (OTU, Chao1, ACE, and Shannon index) of fungal taxa in rumen fluid were clearly higher than parameters in the sweet sorghum straw and the 5 experimental groups (CK, R1 through R7), while the Simpson index was lower than the latter 5 groups. Overall, a decrease in the alpha-diversity parameters of the fungal community was observed with an increase in ensiling time from 30 to 60 days. Among these indices, the OTU, Chao1, and ACE index of the 5 experimental groups (CK, R1 through R7) decreased remarkably. The Shannon indices of 3 groups (CK, R5, and R7) decreased noticeably, while the other 2 groups (R1 and R3) increased with the extension of ensiling time. In contrast, the groups of CK, R5, and R7 increased and another group of R1 and R3 decreased in the Simpson index, displaying a completely different trend.
On the 30th day, indices (OTU, Chao1, and ACE) of 4 experimental groups (R1 through R7) except for the CK group were clearly higher than the raw material sweet sorghum (SS) group. However, the 3 index values were inconsistent when ensiled for 60 days because of the reduction in fungi richness over time. In contrast, the Shannon index of the 5 groups (CK, R1 through R7) was remarkably lower than the SS group, while the Simpson index was remarkably higher. When compared with the CK group, most of the index values (OTU, Chao1, ACE, and Shannon) of the 4 groups (R1 through R7), either 30th or 60th day, were noticeably higher, except for the Shannon index of R3 on the 30th day. Meanwhile, their Simpson index was oppositely lower than the CK group.
In conclusion, with a supplementation of the rumen fluid, the OTU, ACE, Chao1, and Shannon indices of the R1 through R7 groups increased, and the Simpson index decreased.
Table 1. Alpha Diversity Parameters of the Materials and Silage with Different Levels of Rumen Fluid Additives*
Various microorganisms are accommodated in the rumen, including prokaryotes and eukaryotes, possessing a strong capability of converting feed stuffs into microbial biomass and fermentation end products (Wang et al. 2021). Many studies have focused on the bacterial community of the rumen fluid and the silage. For instance, three prokaryotic phyla (Proteobacteria, Firmicutes, and Bacteroidetes) inside the rumen from different age groups were identified as the most predominant, using the 16S rRNA gene sequencing (Wang et al. 2021). Other than the well-known bacteria and protozoa, anaerobic fungi were reported to contribute up to 8% of the microbial mass in rumen and were proposed to significantly engage in decomposition of plant biomass such as CL, HC, and xylans (Kameshwar et al. 2019; Rabee et al. 2019).
Initial ensiling microbiota are composed of natural microbes of forage plant and contaminants (Ávila and Carvalho 2020). In this research, the alpha-diversity of fungi from rumen fluid was much higher than sweet sorghum silage groups, indicating that the rumen fluid can be used as microbial source for bioaugmentation of ensiling. A higher level of value, 1,398 OTUs for fungi in Tibetan goats and sheep rumens, was observed by ITS rRNA gene sequencing (Langda et al. 2020). It was also reported that fungal number of rumen fluid varied greatly ranging between 103 and 106 cells/mL in different studies, with a total of 18 genera, all belonging to the early-branching phylum Neocallimastigomycota (Hagen et al. 2021).
Ensiling time seemed to negatively affect the fungal richness in the present study. It was reviewed that the diversity of moulds changed during the fermentation of silage, resulting in a loss of nutrients and potential contamination by mycotoxins (Ávila and Carvalho 2020). Both fungi and bacteria displayed a significant decreasing trend in the abundance and diversity after 3 days of fermentation, based on the Shannon, Simpson, and Chao1 indices of each sample (Yu et al. 2020). This alteration probably can be explained by the exhaustion of available nutrients, reduction of oxygen and pH value, and formation of other antimicrobial metabolites.
Microbial diversity means the number and abundance of distinct types of microorganisms in a particular ecosystem (Fassarella et al. 2021). The high gut microbial diversity is considered beneficial for human or animal health, for it favors microbe–microbe and host–microbe interactions, while the reduced diversity usually results from the loss of key species essential to keep the equilibrium of the ecosystem, providing opportunities for the enrichment of undesirable microbes, which then potentially impacts microbial resilience and results in disease (Fassarella et al. 2021). A higher richness and diversity of gut microbes enable a more efficient use of nutrient resources (Plaizier et al. 2018). Yaks possessing high diversity of rumen microbes displayed a higher ability to use high-fiber herbage to help them adapt to high-altitude habitats (Fan et al. 2020). Thus, high microbial diversity is closely related to the sustainability and productivity of numerous ecosystems (Fan et al. 2020).
Taxonomic identification and abundance analysis of fungi in rumen fluid
The rumen fluid communities were assigned to 5 phyla including more than 9 genera (Fig. 1). Ascomycota was the most predominant phylum, with a relative abundance of 91.23% (Fig. 1a). The other 4 phyla consisted of Mucoromycota (3.75%), Basidiomycota (2.63%), Mortierellomycota (1.85%), and Neocallimastigomycota (0.55%). At the genus level (Fig. 1b), Penicillium comprised the predominant fungi, with a relative abundance of 62.59%. The other genera included Aspergillus (9.21%), Rhizopus (3.73%), Thermomyces (2.87%), Pichia (2.36%), Mortierella (1.85%), Myceliophthora (1.29%), Candida (1.18%), Cladosporium (1.14%), and other genera (13.78%).
The main phyla and genera observed in this research in rumen fluid of beef heifers and ensilaged samples have also been identified in other ruminants or silages.
Neocallimastigomycota (51.6% in goats, 77.6% in sheep) and Ascomycota (36.0% in goats, 18.0% in sheep) were established as the core fungi in rumens by ITS rRNA gene sequencing (Langda et al. 2020). The two most abundant anaerobic fungi Neocallimastix and Cyllamyces, possessing cellulolytic and xylanolytic enzymes, were investigated in the dromedary camel gut by ITS1 sequencing (Rabee et al. 2019). Three yeasts (Candida sp., Pichia sp., and Trichosporon sp.) and four spoilage molds (Fusarium sp., Aspergillus sp., Muco sp. and Penicillium sp.) were established as the predominant fungi during 60 days fermentation of elephant grass (Vu et al. 2019). Two most abundant phyla, Ascomycota and Basidiomycota, were found in the ensiling of sugarcane tops at the 60th day, and Pichia genus became dominant in the later stage of silage, probably because Pichia can initiate the aerobic deterioration of silages in the later fermentation process (Zhang et al. 2019).
Fig. 1. The relative abundance of fungi community of rumen fluid (a) at the phylum level and (b) at the genera level
Fungi community dynamics during ensiling
The evolution of fungal communities of raw sweet sorghum silages is shown in Fig. 2. The phylum levels of fungi during ensiling are shown in Fig. 2a. Microbiota in the whole ensiling period (60 days) displayed dynamic changes in phylum composition and relative abundance. Five main fungal phyla (Anthophyta, Ascomycota, Basidiomycota, Mortierellomycota, and Mucoromycota) were discovered. The sweet sorghum straw, as the basic materials for fermentation, comprised primarily 3 phylum levels of fungi, composed of Anthophyta, Ascomycota, and Basidiomycota, with a relative abundance of 53.83%, 17.53%, and 28.62%, respectively. With the prolongation of storage time from 0 days (SS group) to 60 days, the population size of Ascomycota increased gradually, while Basidiomycota and Anthophyta showed a downward trend, except the groups of CK and R3 during the 30th to 60th day. Clearly, the fungal communities of groups with rumen fluid added during ensiling were dominated by Ascomycota, with a relative abundance of more than 90%, noticeably higher than SS and CK groups. When ensiled for 60 days, the relative abundance of Ascomycota reached the highest value of 99.66% in the R7 group, while Basidiomycota and Anthophyta displayed a relative abundance of less than 10% in treatment groups (R1 to R7), remarkably lower than in the SS and CK groups. The two lowest levels of relative abundance were Mortierellomycota and Mucoromycota, with less than 1% during the whole storage period.
More detailed fungal compositions at the genus level of the SS and silage groups are shown in Fig. 2b. The dominant genera of the SS group consisted of Cladosporium, Papiliotrema, Hannaella, Rhodotorula, and Vishniacozyma, with the relative abundance of 22.09%, 18.09%, 17.54%, 14.27%, and 12.48%, respectively. In addition, a small number of fungal genera comprised of Pichia, Alternaria, and Mycosphaerella were also detected in the SS group, with the relative abundance of lower than 10%.
Diversity of fungal genera was also found in the CK groups. The predominance on the 30th day consisted of Candida (22.50%), Wickerhamomyces (22.25%), Penicillium (17.05%), Hannaella (12.82%), and Schizophyllum (10.40%). Moreover, a small proportion of fungal population was found with a relative abundance of less than 5%. After 60 days of ensiling, three dominant genera, Schizophyllum (73.90%), Wickerhamomyces (11.83%), and Candida (7.63%), were found in the CK group, while the relative abundance of any other genus was less than 3%, such as Penicillium, Pichia, and other genera.
In treatment groups (R1 through R7) supplemented with rumen fluid, Pichia was established as a dominant genus. With the increase of addition amount of rumen fluid, Pichia displayed a trend of first increasing and then decreasing in the relative abundance, with 56.04% (R1), 71.41% (R3), 93.27% (R5), and 47.01% (R7) on the 30th day, and with 94.39% (R1), 99.23% (R3), 58.28% (R5), and 45.21% (R7) on the 60th day.
Although Penicillium accounted for 42.91% in group R1 after being ensiled of 30 days, it was not found in R3, R5, or R7. Meanwhile, the relative abundance of Candida in R1, R3, and R5 groups was lower than 3%, but increased to 51.64% in group R7, remarkably higher than SS and CK groups.
When the ensiling time lasted for 60 days, the relative abundance of Penicillium, Candida, and Schizophyllum was less than 1% in the R1 and R3 groups. With the increase of rumen fluid addition and the trend variation in the Pichia quantity, Schizophyllum and Candida became another 2 dominances besides Pichia in group R5, with the relative abundance of 24.29% and 11.18%, respectively. In group R7, the two genera changed to 0.02% and 48.13%, respectively.
In conclusion, when compared with SS and CK groups, the use of rumen fluid remarkably increased the relative abundances of Pichia and accelerated the reduction of Schizophyllum and Penicillium in silages.
Fig. 2. The fungal communities and relative abundance at the (a) phylum and (b) genus levels of raw and ensiled sweet sorghum; SS = raw sweet sorghum; CK = control group without rumen fluid addition; R1, R3, R5, and R7 = rumen fluid supplemented with dosages of 1, 3, 5, and 7 g/100 g of wet raw sweet sorghum, respectively. The numbers 30 and 60 indicate 30 and 60 days of silage, respectively.
Saccharomyces and Candida, as well as Pichia, are regarded as the main initiators of the aerobic spoilage of silage, because of their lactate-assimilating, facultative anaerobic, and acid tolerant characteristics (Ávila and Carvalho 2020). It is probably for these above abilities and the adaptability to the ensiling environment that Pichia in the current study became a predominant fungus in silage following the bioaugmentation, even though its abundance was much lower than others in rumen fluid.
Penicillium and Aspergillus are generally considered as toxigenic fungi during ensiling (Mostrom and Jacobsen 2020). However, both fungi were found displaying strong cellulolytic activities, and producing all three components of cellulase (endocellulase, exocellulase, and β-glucosidase) (Vaishnav et al. 2018). Aspergillus spp. has been regarded as the most frequent aerobic fungus in rumen fluid from beef heifers of different ages and sex in previous reports (Abrão et al. 2014; Liu et al. 2020). Thus, the two most predominant fungi in rumen fluid in the present research, Penicillium and Aspergillus, perhaps positively promoted CL decomposition of silage, and would be desirable when equipped with the detection of mycotoxins during the ensiling process. But with the addition of the rumen fluid, a reduction of Penicillium and Schizophyllum in silage was found, probably due to inhibition by the acidic and anaerobic environment resulting from LAB metabolism (Yu et al. 2020).
Correlations between fungus communities and chemical compositions/fermentative characteristics
Figure 3a through 3f shows the heat map of the correlation between the fungal communities and the fermentation quality and chemical index of ensiled sweet sorghum on the 30th and 60th days of storage.
As shown in Figs. 3a and 3d, the fermentation characteristics of silage was assessed from aspects of contents of LA, AA, AN, and pH value. When ensiled for 30 days (Fig. 3a), the abundance of Cladosporium was significantly negatively correlated with the contents of LA and AA, with the Spearman correlation coefficient (rs) of -0.943 (P < 0.01) and -0.886 (P < 0.05), respectively. Moreover, three other genera containing Hannaella, Alternaria, and Vishniacozyma, displayed significance of negative correlation with AA contents, with the rs = -0.942 (P < 0.01), -0.886 (P < 0.05), and -0.885 (P < 0.05), respectively. Correspondingly, pH values of silage samples showed significantly positive correlation with the above three genera, meaning that growth of these microbes can cause the pH increase. After ensiled for 60 days (Fig. 3d), the abundance of Alternaria (rs = 0.801, P < 0.01) and Saccharomyces (rs = 0.804, P < 0.01) showed significant negative correlation with the change of AN content. A significant positive correlation between the Pichia abundance and the contents of LA and AA was found, with rs = 0.766 (P < 0.05) and rs = 0.679 (P < 0.05), respectively. Correspondingly, the Pichia genus was negatively correlated with the pH value (rs = 0.818, P < 0.01). In conclusion, Pichia was conductive to the ensiled quality, while the Hannaella and Vishniacozyma genera appeared to be unfavorable for the ensiling process.
Furthermore, the amounts of dry matter (DM) and crude protein (CP) of silages stored for 30 days (Fig. 3b) displayed a significant positive correlation with survival of Pichia and Cladosporium, while showing a significant negative correlation with Alternaria growth. Water-soluble carbohydrates (WSC) content was significantly positively correlated with Cladosporium (rs = 0.683, P < 0.05) on the 30th day, while significantly negatively correlated with Mucor (rs = -0.758, P < 0.05) on the 60th day (Fig. 3e). Thus, Pichia and Cladosporium genera were beneficial to the nutrient preservation during ensiling from correlation analysis.
Correlations between fungal flora and fiber components are displayed in Figs. 3c and 3f, including NDF, ADF, and acid detergent lignin (ADL), along with the hemicellulose (HC) in Figs. 3b and 3e.
Fig. 3. Correlations of fungus diversity at genus level with: (a, b) CP, DM, WSC, and HC; (c, d) ADF, NDF, ADL, and CL; and (e, f) pH, LA, AA, and AN after ensiling for 30 days or 60 days. The corresponding value of the heat map is the Spearman correlation coefficient rs, which ranges between −1 and 1. Furthermore, rs < 0 indicates a negative correlation (blue), and rs > 0 indicates a positive correlation (red). * P < 0.05, ** P < 0.01, and *** P < 0.001.
Pichia and Cladosporium were significantly positively correlated with the contents of NDF, ADF, and ADL, when ensiled for 30 days. Meanwhile, Hannaella displayed a significant negative correlation with contents of ADF and NDF. With the increase of ensiling time, Vishniacozyma was significantly positively correlated with the contents of CL and ADF on the 60th day, with rs = -0.728 (P < 0.05) and rs = -0.815 (P < 0.05), respectively.
The other fungal flora, during the whole ensiling process (30 and 60 days), revealed no significant correlation with the fermentation quality and chemical indices of sweet sorghum silage.
It is universally known that rumen microbiota in ruminants is vital for sustaining good rumen ecology, health, and productivity (Fan et al. 2020). Similarly, certain microbial groups could directly or indirectly affect the quality of forage during ensiling. Generally, bacteria containing LAB act as an important role in the fermentative conversion of forage substrate, for instance, Prevotella spp. and Selenomonas ruminantium participate in the degradation of lactate and other oligosaccharides, Ruminococcus albus and Ruminococcus flavefaciens are usually considered as cellulolytic bacteria (Wang et al. 2021). While yeasts and moulds are considered as the most important putrefactive organisms in silage, for their growth could lead to loss of nutrients, mycotoxin production, and reduction in palatability, causing losses in animal performance (Vu et al. 2019).
However, various hydrolases secreted by fungi (cellulases, xylanases, mannases, esterases, glucosidases, and glucanases) and the rhizoid system could enable them to penetrate plant cell walls and mechanically degrade the plant substrate (Fliegerova et al. 2015). Some studies suggest that contribution of the anaerobic gut fungi to fermentation can greatly exceed bacteria (Lee et al. 2000). When anaerobic rumen fungi were removed from the rumen, feed intake by sheep decreased but was restored after the anaerobic fungus Neocallimastix sp. re-inoculated back (Gordon and Phillips 1993).
Information about the role and microbial interactions of fungi during ensiling is not enough. In the rumen, anaerobic fungi provide an increased surface area for bacterial colonization and available nutrient substance by both the mechanical and enzymatic degradation of plant material (Gruninger et al. 2014). Supplementation of Saccharomyces cerevisiae increased the OTU richness related to fibrolytic bacteria Fibrobacter succinogenes and Prevotella (AlZahal et al. 2017). This relationship in silage process may benefit the production of LA and AA by LAB, which consist of homofermentative and heterofermentative groups. However, the metabolites of bacteria probably negatively affect the growth of some fungi under anaerobic environment, except for the genera of acid-tolerant components such as Pichia yeast. As a facultative anaerobe, yeasts can produce substances such as CO2 and ethanol using sugars. In addition, yeasts can increase the protein content in silage, but also lead to the aerobic spoilage, leading to the reduction of the quality of silage (Drouin et al. 2021). Although yeasts and moulds are generally considered undesirable in silage, Pichia as well as Cladosporium in this study showed a positive role in maintaining the nutrients during ensiling. This phenomenon may be related to the silage stage, because during the 60 days period the silage was not opened, and aerobic exposure was not involved. Of course, further mechanisms will be studied in the future.
The rumen fluid was collected as a by-product from cattle slaughter plants, and less than 10 L of rumen fluid can be obtained from one cow or bull, which probably limits the industrial application of rumen fluid in pretreatment for silage, although the addition rate of rumen fluid into silage was relatively low. On the other hand, the application of rumen fluid in silage pretreatment provides a good prospect for the production of high-quality silage, which have been tested by various researchers (Krause et al. 2013; Wang et al. 2019; Ren et al. 2021). Among the microbes in rumen fluid, the desirable fungi as well as LAB, playing a key role during digestion of ruminants and silage fermentation, will be identified and cultured in future. Furthermore, these isolates may be used as adjunct inoculants for the improvement of forage quality.
CONCLUSIONS
- The rumen fluid additive can effectively reduce the reproduction of adverse fungi. The microbiota of rumen fluid, as well as silage, is a complex mix of fungi, bacteria, and other microbes. Interaction of microbes between the two niches greatly affected the quality of silage. Bioaugmentation of sweet sorghum ensiling with rumen fluid and the present research on diversity and dynamics of fungi community will be favorable to the improvement of ensiling fermentation in the future.
- Rumen fluid supplementation exerted a remarkable influence on the fungal diversity of sweet sorghum silage. A significant increase in the relative abundance of Pichia and a decrease in Schizophyllum and Penicillium were observed, following the supplementation of rumen fluid, which, however, was high in the abundance of Penicillium and Aspergillus.
- Correlation analysis suggested that Pichia was significantly positively correlated with the fermentation quality of silage, while Hannaella and Vishniacozyma were significantly negatively correlated with the ensiling characteristics.
ACKNOWLEDGMENTS
This work was financially supported by the Key Project of Natural Science Foundation of Gansu Province (Grant No. 21JR7RA203), the China Postdoctoral Science Foundation (Grant No. 2019T120961), West Light Foundation of Chinese Academy of Sciences (Grant No. 2018XBZG XBQNXZ A), Red Willow Distinguished Young Talents (Grant No. JQ2020), and First-class Discipline Project of Lanzhou University of Technology (Grant No. 0807J1).
Ethics Statement
All animal experiments were performed following the management guidelines of the Gansu Ethical Review Committee (License No. SYXK-GAN-2014-003). All animals used were humanely bled and euthanized appropriately for experimental purposes.
Abrão, F. O., Duarte, E. R., Freitas, C. E. S., Vieira, E. A., Geraseev, L. C., Da Silva-Hughes, A. F., Rosa, C. A., and Rodrigues, N. M. (2014). “Characterization of fungi from ruminal fluid of beef cattle with different ages and raised in tropical lignified pastures,” Current Microbiology 69(5), 649-659. DOI: 10.1007/s00284-014-0633-5
AlZahal, O., Li, F., Guan, L. L., Walker, N. D., and McBride, B. W. (2017). “Factors influencing ruminal bacterial community diversity and composition and microbial fibrolytic enzyme abundance in lactating dairy cows with a focus on the role of active dry yeast,” Journal of Dairy Science 100(6), 4377-4393. DOI: 10.3168/jds.2016-11473
Ambye-Jensen, M., Johansen, K. S., Didion, T., Kádár, Z., Schmidt, J. E., and Meyer, A. S. (2013). “Ensiling as biological pretreatment of grass (Festulolium Hykor): The effect of composition, dry matter, and inocula on cellulose convertibility,” Biomass and Bioenergy 58, 303-312. DOI: 10.1016/j.biombioe.2013.08.015
Ávila, C. L. S., and Carvalho, B. F. (2020). “Silage fermentation—Updates focusing on the performance of micro-organisms,” Journal of Applied Microbiology 128(4), 966-984. DOI: 10.1111/jam.14450
del Palacio, A., Mionetto, A., Bettucci, L., and Pan, D. (2016). “Evolution of fungal population and mycotoxins in sorghum silage,” Food Additives & Contaminants: Part A 33(12), 1864-1872. DOI: 10.1080/19440049.2016.1244732
Doyle, N., Mbandlwa, P., Kelly, W. J., Attwood, G., Li, Y., Ross, R. P., Stanton, C., and Leahy, S. (2019). “Use of lactic acid bacteria to reduce methane production in ruminants, a critical review,” Frontiers in Microbiology 10, article no. 2207. DOI: 10.3389/fmicb.2019.02207
Drouin, P., and Ferrero, F. (2020). “Testing selectivity of bacterial and fungal culture media compared to original silage samples using next generation sequencing,” Journal of Microbiological Methods 179, article ID 106088. DOI: 10.1016/j.mimet.2020.106088
Drouin, P., Tremblay, J., Renaud, J., and Apper, E. (2021). “Microbiota succession during aerobic stability of maize silage inoculated with Lentilactobacillus buchneri NCIMB 40788 and Lentilactobacillus hilgardii CNCM-I-4785,” MicrobiologyOpen 10(1), article ID e1153. DOI: 10.1002/mbo3.1153
Edgar, R. C. (2013). “UPARSE: Highly accurate OTU sequences from microbial amplicon reads,” Nature Methods 10(10), 996-998. DOI: 10.1038/nmeth.2604
Fan, Q., Wanapat, M., Yan, T., and Hou, F. (2020). “Altitude influences microbial diversity and herbage fermentation in the rumen of yaks,” BMC Microbiology 20(1), article Number 370. DOI: 10.1186/s12866-020-02054-5
Fassarella, M., Blaak, E. E., Penders, J., Nauta, A., Smidt, H., and Zoetendal, E. G. (2021). “Gut microbiome stability and resilience: Elucidating the response to perturbations in order to modulate gut health,” Gut 70(3), 595-605. DOI: 10.1136/gutjnl-2020-321747
Fliegerova, K., Kaerger, K., Kirk, P., and Voigt, K. (2015). “Rumen fungi,” in: Rumen Microbiology: From Evolution to Revolution, A. K. Puniya, R. Singh, and D. N. Kamra (eds.), Springer, New Delhi, India, pp. 97-112. DOI: 10.1007/978-81-322-2401-3_7
Gordon, G. L. R., and Phillips, M. W. (1993). “Removal of anaerobic fungi from the rumen of sheep by chemical treatment and the effect on feed consumption and in vivo fibre digestion,” Letters in Applied Microbiology 17(5), 220-223. DOI: 10.1111/j.1472-765X.1993.tb01451.x
Gruninger, R. J., Puniya, A. K., Callaghan, T. M., Edwards, J. E., Youssef, N., Dagar, S. S., Fliegerova, K., Griffith, G. W., Forster, R., Tsang, A., et al. (2014). “Anaerobic fungi (phylum Neocallimastigomycota): Advances in understanding their taxonomy, life cycle, ecology, role and biotechnological potential,” FEMS Microbiology Ecology 90(1), 1-17. DOI: 10.1111/1574-6941.12383
Guo, G., Shen, C., Liu, Q., Zhang, S., Shao, T., Wang, C., Wang, Y., Xu, Q., and Huo, W. (2020). “The effect of lactic acid bacteria inoculums on in vitro rumen fermentation, methane production, ruminal cellulolytic bacteria populations and cellulase activities of corn stover silage,” Journal of Integrative Agriculture 19(3), 838-847. DOI: 10.1016/S2095-3119(19)62707-3
Hagen, L. H., Brooke, C. G., Shaw, C. A., Norbeck, A. D., Piao, H., Arntzen, M. Ø., Olson, H. M., Copeland, A., Isern, N., Shukla, A., et al. (2021). “Proteome specialization of anaerobic fungi during ruminal degradation of recalcitrant plant fiber,” The ISME Journal 15(2), 421-434. DOI: 10.1038/s41396-020-00769-x
Huang, D., Xu, Y., Lei, F., Yu, X., Ouyang, Z., Chen, Y., Jia, H., and Guo, X. (2021). “Degradation of polyethylene plastic in soil and effects on microbial community composition,” Journal of Hazardous Materials 416, article ID 126173. DOI: 10.1016/j.jhazmat.2021.126173
Kameshwar, A. K. S., Ramos, L. P., and Qin, W. (2019). “Metadata analysis approaches for understanding and improving the functional involvement of rumen microbial consortium in digestion and metabolism of plant biomass,” Journal of Genomics 7, 31-45. DOI: 10.7150/jgen.32164
Kharazian, Z. A., Jouzani, G. S., Aghdasi, M., Khorvash, M., Zamani, M., and Mohammadzadeh, H. (2017). “Biocontrol potential of Lactobacillus strains isolated from corn silages against some plant pathogenic fungi,” Biological Control 110, 33-43. DOI: 10.1016/j.biocontrol.2017.04.004
Krause, D. O., Nagaraja, T. G., Wright, A. D., and Callaway, T. R. (2013). “Board-invited review: Rumen microbiology: Leading the way in microbial ecology,” Journal of Animal Science 91(1), 331-341. DOI: 10.2527/jas.2012-5567
Kung, L., Shaver, R. D., Grant, R. J., and Schmidt, R. J. (2018) “Silage review: Interpretation of chemical, microbial, and organoleptic components of silages,” Journal of Dairy Science 101(5), 4020-4033. DOI: 10.3168/jds.2017-13909
Langda, S., Zhang, C., Zhang, K., Gui, B., Ji, D., Deji, C., Cuoji, A., Wang, X., and Wu, Y. (2020). “Diversity and composition of rumen bacteria, fungi, and protozoa in goats and sheep living in the same high-altitude pasture,” Animals (Basel) 10(2), Article Number 186. DOI: 10.3390/ani10020186
Lee, S. S., Ha, J. K., and Cheng, K. (2000). “Relative contributions of bacteria, protozoa, and fungi to in vitro degradation of orchard grass cell walls and their interactions,” Applied and Environmental Microbiology 66(9), 3807-3813. DOI: 10.1128/AEM.66.9.3807-3813.2000
Li, F., Ding, Z., Ke, W., Xu, D., Zhang, P., Bai, J., Mudassar, S., Muhammad, I., and Guo, X. (2019). “Ferulic acid esterase-producing lactic acid bacteria and cellulase pretreatments of corn stalk silage at two different temperatures: Ensiling characteristics, carbohydrates composition and enzymatic saccharification,” Bioresource Technology 282, 211-221. DOI: 10.1016/j.biortech.2019.03.022
Li, S., Deng, Y., Wang, Z., Zhang, Z., Kong, X., Zhou, W., Yi, Y., and Qu, Y. (2020). “Exploring the accuracy of amplicon-based internal transcribed spacer markers for a fungal community,” Molecular Ecology Resources 20(1), 170-184. DOI: 10.1111/1755-0998.13097
Liu, B., Huan, H., Gu, H., Xu, N., Shen, Q., and Ding, C. (2018). “Dynamics of a microbial community during ensiling and upon aerobic exposure in lactic acid bacteria inoculation-treated and untreated barley silages,” Bioresource Technology 273, 212-219. DOI: 10.1016/j.biortech.2018.10.041
Liu, B., Yang, Z., Huan, H., Gu, H., Xu, N., and Ding, C. (2020). “Impact of molasses and microbial inoculants on fermentation quality, aerobic stability, and bacterial and fungal microbiomes of barley silage,” Scientific Reports 10(1), article no. 5342. DOI: 10.1038/s41598-020-62290-7
Mackie, R., Aminov, R., White, B. A., and Mcsweeney, C. (2000). “Molecular ecology and diversity in gut microbial ecosystems,” in: Ruminant Physiology: Digestion, Metabolism, Growth and Reproduction, P. B. Gronje (ed.), CAB International, Wallingford, UK, pp. 61-77. DOI: 10.1079/9780851994635.0061
Mostrom, M. S., and Jacobsen, B. J. (2020). “Ruminant mycotoxicosis: An update,” Veterinary Clinics of North America: Food Animal Practice 36(3), 745-774. DOI: 10.1016/j.cvfa.2020.08.011
Nkosi, B. D., Vadlani, P. V., Brijwani, K., Nanjunda, A., and Meeske, R. (2012). “Effects of bacterial inoculants and an enzyme on the fermentation quality and aerobic stability of ensiled whole-crop sweet sorghum,” South African Journal of Animal Science 42(3), 232-240. DOI: 10.4314/sajas.v42i3.4
Panasiuk, L., Jedziniak, P., Pietruszka, K., Piatkowska, M., and Bocian, L. (2019). “Frequency and levels of regulated and emerging mycotoxins in silage in Poland,” Mycotoxin Research 35(1), 17-25. DOI: 10.1007/s12550-018-0327-0
Plaizier, J. C., Danesh, M. M., Derakhshani, H., Golder, H., Khafipour, E., Kleen, J. L., Lean, I., Loor, J., Penner, G., and Zebeli, Q. (2018). “Review: Enhancing gastrointestinal health in dairy cows,” Animal 12(s2), s399-s418. DOI: 10.1017/S1751731118001921
Rabee, A. E., Forster, R. J., Elekwachi, C. O., Kewan, K. Z., Sabra, E. A., Shawket, S. M., Mahrous, H. A., and Khamiss, O. A. (2019). “Community structure and fibrolytic activities of anaerobic rumen fungi in dromedary camels,” Journal of Basic Microbiology 59(1), 101-110. DOI: 10.1002/jobm.201800323
Ren, H., Sun, W., Yan, Z., Zhang, Y., Wang, Z., Song, B., Zheng, Y., and Li, J. (2021). “Bioaugmentation of sweet sorghum ensiling with rumen fluid: Fermentation characteristics, chemical composition, microbial community, and enzymatic digestibility of silages,” Journal of Cleaner Production 294, Article ID 126308. DOI: 10.1016/j.jclepro.2021.126308
Romero, J. J., Zhao, Y., Balseca-Paredes, M. A., Tiezzi, F., Gutierrez-Rodriguez, E., and Castillo, M. S. (2017). “Laboratory silo type and inoculation effects on nutritional composition, fermentation, and bacterial and fungal communities of oat silage,” Journal of Dairy Science 100(3), 1812-1828. DOI: 10.3168/jds.2016-11642
Sirohi, S. K., Singh, N., Dagar, S. S., and Puniya, A. K. (2012). “Molecular tools for deciphering the microbial community structure and diversity in rumen ecosystem,” Applied Microbiology and Biotechnology 95(5), 1135-1154. DOI: 10.1007/s00253-012-4262-2
Vaishnav, N., Singh, A., Adsul, M., Dixit, P., Sandhu, S. K., Mathur, A., Puri, S. K., and Singhania, R. R. (2018). “Penicillium: The next emerging champion for cellulase production,” Bioresource Technology Reports 2, 131-140. DOI: 10.1016/j.biteb.2018.04.003
Vu, V. H., Li, X., Wang, M., Liu, R., Zhang, G., Liu, W., Xia, B., and Sun, Q. (2019). “Dynamics of fungal community during silage fermentation of elephant grass (Pennisetum purpureum) produced in northern Vietnam,” Asian-Australasian Journal of Animal Sciences 32(7), 996-1006. DOI: 10.5713/ajas.18.0708
Wang, D., Zhao, C., Liu, S., Zhang, T., Yao, J., and Cao, Y. (2019). “Effects of Piromyces sp. CN6 CGMCC 14449 on fermentation quality, nutrient composition and the in vitro degradation rate of whole crop maize silage,” AMB Express 9(1), article no. 121. DOI: 10.1186/s13568-019-0846-x
Wang, Z., Yang, D. S., Li, X. Y., Yu, Y. N., Yong, L. Y., Zhang, P. H., He, J. H., Shen, W. J., Wan, F. C., Feng, B. L., et al. (2021). “Modulation of rumen fermentation and microbial community through increasing dietary cation-anion difference in Chinese Holstein dairy cows under heat stress conditions,” Journal of Applied Microbiology 130(3), 722-735. DOI: 10.1111/jam.14812
Wang, Q., Wang, R., Wang, C., Dong, W., Zhang, Z., Zhao, L., and Zhang, X. (2022). “Effects of cellulase and Lactobacillus plantarum on fermentation quality, chemical composition, and microbial community of mixed silage of whole-plant corn and peanut vines,” Applied Biochemistry and Biotechnology. DOI: 10.1007/s12010-022-03821-y
Yu, J., Hou, Q., Li, W., Huang, W., Mo, L., Yao, C., An, X., Sun, Z., and Wei, H. (2020). “Profiling of the viable bacterial and fungal microbiota in fermented feeds using single-molecule real-time sequencing,” Journal of Animal Science 98(2), 1-10. DOI: 10.1093/jas/skaa029
Yue, Z., Li, W., and Yu, H. (2013). “Application of rumen microorganisms for anaerobic bioconversion of lignocellulosic biomass,” Bioresource Technology 128, 738-744. DOI: 10.1016/j.biortech.2012.11.073
Zhang, H., Zhang, P., Ye, J., Wu, Y., Fang, W., Gou, X., and Zeng, G. (2016). “Improvement of methane production from rice straw with rumen fluid pretreatment: A feasibility study,” International Biodeterioration & Biodegradation 113, 9-16. DOI: 10.1016/j.ibiod.2016.03.022
Zhang, L., Zhou, X., Gu, Q., Liang, M., Mu, S., Zhou, B., Huang, F., Lin, B., and Zou, C. (2019). “Analysis of the correlation between bacteria and fungi in sugarcane tops silage prior to and after aerobic exposure,” Bioresource Technology 291, article ID 121835. DOI: 10.1016/j.biortech.2019.121835
Article submitted: December 23, 2021; Peer review completed: March 20, 2022; Revised version received and accepted: April 2, 2022; Published: April 6, 2022.
DOI: 10.15376/biores.17.2.2977-2996
