Abstract
Recently, xylanase has attracted attention in the pulp and paper industry because of its bleach-boosting properties, which reduces chemical consumption during the bleaching process. Four fungi species, namely Penicillium oxalicum T3.3, Aspergillus niger ATCC 6275, Colletotrichum gloeosporioides,and Pycnoporus sanguineus, were studied to select which fungi can best produce cellulase-poor xylanase using rice straw as a substrate. The fungi were screened for xylanase production based on the clear zone formation on a malt extract agar (MEA) plate containing xylan. P. oxalicum T3.3 and A. niger ATCC 6275 showed greater clear zone formation on the agar plate than the other two studied fungi. These fungi were grown in a liquid medium containing rice straw as the substrate. P. oxalicum T3.3 showed the highest xylanase activity (65.89 U/mL) with lowest carboxymethyl cellulase (CMCase) (1.88 U/mL) and filter paperase (FPase) activity (0.16 U/mL) after four days of fermentation at 30 °C. P. oxalicum T3.3 produced higher xylanase activity when rice straw was used as a substrate compared to commercial xylan. Thus, cellulase-poor xylanase produced from P. oxalicum T3.3 has high potential for biobleaching application in the pulp and paper industry in terms of technical and biological performance and economical aspects.
Download PDF
Full Article
Selection of Potential Fungi for Production of Cellulase-Poor Xylanase from Rice Straw
Nur Idayu Zahari,a Umi Kalsom Md Shah,a,b,c,* Ainun Zuriyati Mohamed Asa’ari,b and Rosfarizan Mohamad a,b,c
Recently, xylanase has attracted attention in the pulp and paper industry because of its bleach-boosting properties, which reduces chemical consumption during the bleaching process. Four fungi species, namely Penicillium oxalicum T3.3, Aspergillus niger ATCC 6275, Colletotrichum gloeosporioides, and Pycnoporus sanguineus, were studied to select which fungi can best produce cellulase-poor xylanase using rice straw as a substrate. The fungi were screened for xylanase production based on the clear zone formation on a malt extract agar (MEA) plate containing xylan. P. oxalicum T3.3 and A. niger ATCC 6275 showed greater clear zone formation on the agar plate than the other two studied fungi. These fungi were grown in a liquid medium containing rice straw as the substrate. P. oxalicum T3.3 showed the highest xylanase activity (65.89 U/mL) with lowest carboxymethyl cellulase (CMCase) (1.88 U/mL) and filter paperase (FPase) activity (0.16 U/mL) after four days of fermentation at 30 °C. P. oxalicum T3.3 produced higher xylanase activity when rice straw was used as a substrate compared to commercial xylan. Thus, cellulase-poor xylanase produced from P. oxalicum T3.3 has high potential for biobleaching application in the pulp and paper industry in terms of technical and biological performance and economical aspects.
Keywords: Xylanase; Penicillium oxalicum T3.3; Rice straw; Biobleaching; Pulp and paper
Contact information: a: Department of Bioprocess Technology, Universiti Putra Malaysia; b: Institute of Tropical Forestry and Forest Products, Universiti Putra Malaysia; c: Bioprocessing and Manufacturing Research Centre, Universiti Putra Malaysia 43400 UPM Serdang, Selangor, Malaysia;
* Corresponding author: umikalsom@upm.edu.my
INTRODUCTION
Growing consumer demand for environmentally safe products and processes has had a major influence on the pulp and paper industry. Like other industries, the pulp and paper industry can significantly impact the environment. The chlorinated organic compounds and absorbable organic halides that are produced during the bleaching are harmful to biological systems (Beg et al. 2000; Ali and Sreekrishnan 2001; Techapun et al. 2003). Thus, to reduce pollution, eco-friendly approaches are often suggested, such as using enzymes. Currently, xylanase enzyme production focuses more on the application of fiber processing in various industries. The enzyme is used to pretreat pulps prior to bleaching, which is known as the biobleaching process (Ghatora et al. 2006). The use of xylanase enzymes enhances the bleaching by removing the region of xylan with high hexenuronic acid content and facilitating the release of lignin from the pulp (Henriksson and Teeri 2009). Therefore, xylanase reduces the usage of chlorine during the bleaching process. In the pulp and paper industry, the removal of lignin is required to produce brighter or whiter pulps. Chromophores, which are located on lignin in all woody or nonwoody plants, causes absorption of visible light by the wood, pulp, or fibres (Koponen 1991; Beg et al. 2001; Sahin and Mantanis 2011; Nagar et al. 2013). Successful biobleaching occurs when the enzyme used is stable and active at both temperature and alkaline conditions (Beg et al. 2000; Gupta et al. 2000). The biobleaching process also requires the use of cellulase-free xylanase or low cellulase enzymes (Nagar et al. 2013). A high level of cellulase activity results in loss of cellulose, degradation of pulp quality, and an increase in effluent treatment cost (Beg et al. 2001). According to Liao et al. (2015), P. oxalicum GZ-2 has different types of xylanase genes, which are xyn10A, xyn11A, xyn10B, and xyn11B that produced different types of xylanase. These xylanase have different properties. But for the biobleaching process, pure xylanase is not required, and this can save costs during production. Besides, the cloning process is an expensive process.
Xylanases are glycosidases, which catalyze the hydrolysis of 1,4-β-D-xylosidic linkages in xylan (Hatanaka 2012). The depolymerisation action of xylanase produces monomeric pentose units, xylobiose, xylotriose, and substituents containing two or four xylose residues that are used by microorganisms as a primary carbon source (McCrae et al. 1994; Latif et al. 2006; Kanimozhi and Nagalakshmi 2014). Xylanases differ in their specificity towards the xylan polymer. The bond selected for hydrolysis depends on the nature of the substrate molecule, including the chain length, the degree of branching, and the presence of substituents (Li et al. 2000). Xylan’s varying and complex structure results in a great diversity of xylanase with varying specificities, primary sequences, and folds (Collins et al. 2005).
From an industrial point of view, filamentous fungi are particularly interesting for xylanase production. This is due to their high production level and extracellular secretion of enzymes, as well as the relative ease of cultivation. Generally, xylanase activity levels from fungal cultures are typically higher than yeast or bacteria (Paloheimo et al. 2003). For fungi producing xylanase, particular attention has been granted to the enzymes produced by Aspergillus, Trichoderma, and Penicillium (Abdel-sater and El-said 2001). Xylanase production that uses a variety of lignocellulosic biomass has been reported by many researchers (Okafor et al. 2007).
Agricultural activities such as crop production, crop harvest, saw milling, and agro-industrial processing produce a large amount of lignocellulosic biomass (Pang and Ibrahim 2005). Accumulation of these wastes without recycling can cause environmental pollution (Abu et al. 2000; Norazlina et al. 2013). Because of the biodegradable properties, these lignocellulosic wastes can be managed and converted into valuable products such as biofuels, chemicals, cheap energy source for fermentation, improved animal feed, and human nutrients (Howard et al. 2003).
In many countries where rice is a major crop, rice straw is a primary agricultural by-product (Sun et al. 2008). According to the United States Department of Agriculture (USDA 2015), rice production in the world was estimated at 481.74 million tonnes in 2015/2016, an increase of 5.62 million tonnes from the previous year. China, which is the largest global producer and exporter of rice, produces around 204.28 million tonnes of rice per year, India produces 152.60 million tonnes, and Indonesia produces 69.04 million tonnes (FAO 2014). According to FAO (2010), Malaysia produces over 2.5 million tonnes of rice. In Malaysia, the ratio of rice harvest to straw is fairly consistent from year to year, and the ratio 1:1 was used to estimate the amount of straw that will be obtained (MADA 2010). Rice straw is mostly composed of cellulose, hemicellulose, and lignin. The major portion of hemicellulose is known as xylan, which represents approximately 20% to 35% of the total dry weight in tropical plant biomass (Levasseur et al. 2005). A molecule of xylan is the major structural heteropolysaccharide in plants consisting of β-1, 4-linked polymers of xylopyranosyl units, and it can be substituted by side groups such as L-arabinose, D-galactose, acetyl, feruloyl, p-coumarouyl, and glucuronic acid residues (Subramaniyan and Prema 2002; Bakri et al. 2010). Previous reports have shown that xylanase is an inducible enzyme. Thus, the production of xylanase enzyme is enhanced when the fungus is grown in a medium containing xylan such as rice straw, wheat straw, acetylated oat spelt xylan, and arabitol, but the level of production varies (Eyzaguirre et al. 1992; Egaña et al. 1996; De Ioannes et al. 2000).
Even though a lot of commercial xylanase products are available at industrial scale, only a few are used in pulp and paper industry such as Cartazyme, EcopulpX-200, Pulpzyme and Xylanase GS 35, which are specialized in bleaching of pulp (Harris and Ramalingam 2010). Xylanase from bacteria might have higher values for application in biobleaching, yet they have low activity compared to fungi. The high activity of crude enzyme is very important when the enzymes are not applied in pure form (Kulkarni et al. 1999; Subramaniyan and Prema 2002). Therefore, the aim of this study was to determine potential fungi that could best produce cellulase poor xylanase using a renewable lignocellulosic biomass, such as rice straw, as an economical substrate.
EXPERIMENTAL
Microorganisms and Culture Condition
The fungi species P. oxalicum T3.3 is an endophytic fungus isolated from dragon fruit stem. Additionally, during this study, P. sanguineus was isolated from wood obtained from the Department of Bioprocess Technology, Faculty of Biotechnology and Biomolecular Sciences, Universiti Putra Malaysia. C. gloeosporioides was obtained from the Strategic Resource Research Centre, Malaysian Agricultural Research and Development Institute (MARDI), while A. niger ATCC 6275 was obtained from the American Type Culture Collection. To prolong the life and expand the number of cells, these fungi were subcultured on potato dextrose agar at 30 °C for seven days and maintained at 4 °C.
Plate Screening for Xylanase Activity on Malt Extract Agar (MEA)
The purpose of primary screening was to obtain fungi that were able to produce xylanase enzyme. Plate screening is a qualitative method used to identify xylanase producers using selective agar plates to produce different sizes of clearing zone (Wong et al. 1988). The four fungal species were screened based on their abilities to produce extracellular xylanase during their growth on enriched malt extract agar (MEA) that contains 0.5 % (w/v) xylan as the carbon source. The fungi were incubated for 3 to 4 days at 28 ± 2 °C (Nakamura 2003). Xylanase producers were detected based on the clear zones of xylan hydrolysis around the colony after the plates were flooded with 0.1% (w/v) aqueous Congo Red for 15 min, followed by washing with 1 M NaCl (Sridevi and Charya 2011). The clear zones were measured and the relative enzyme activity of each isolate was determined. The relative enzyme activity was determined by dividing the diameter of the clearing zone (mm) with the diameter of mycelia growth (mm) (Adesina and Onilude 2013). The fungi species that produced clear zones around the colonies were selected for the next experiment.
Enzyme Production by Submerged Fermentation
The selected fungi were subjected to submerged fermentation to quantitatively estimate the xylanase enzyme production. Three discs (1 cm in diameter) of mycelia from seven-day-old cultures of P. oxalicum T3.3 were inoculated in triplicate in 250-mL Erlenmeyer flasks containing 100 mL of Mandel’s medium. The Mandel’s media was composed of (NH4)2SO4, 1.40 g; KH2PO4, 2.00 g; urea, 0.63 g; MgSO4.7H2O, 0.30 g; trace element, 1.00 mL; CaCl2, 0.30 g; yeast extract, 0.75 g; Tween 80, 2.00 mL; and rice straw, 1.00 g, per litre of distilled water (Mandels et al. 1974). The flasks were shaken for six days at 30 °C on a rotary shaker at 150 rpm. Samples were periodically drawn every 24 h. Culture filtrates from the fermentation broth were centrifuged at 10,000 rpm for 10 min at room temperature, and the supernatants were tested for enzyme activity assay (Bakri et al. 2008). In this experiment, A. niger ATCC 6275 and commercial xylan from beechwood were used as the control model.
Quantitative Assay for Xylanase, Cellulase Enzyme, and Protein
Xylanase assay
Xylanase enzyme activity was assayed by measuring the reducing sugar liberated from 1% (w/v) beechwood xylan (Sigma-Aldrich Co., St. Louis, MO) dissolved in 50 mM sodium citrate buffer (pH 5.0). The reaction mixture (composed of 1.8 mL xylan and 0.2 mL crude enzyme) was incubated in a water bath at 40 °C for 30 min. The reaction was stopped by adding 3 mL of 3,5-dinitrosalicylic acid (DNS) and incubated in boiling water for 15 min. After cooling, the sample was read at 575 nm using a spectrophotometer (UvLine 9400 by Secomman, France). The amount of reducing sugar was measured using xylose as the standard. One unit of enzyme activity is defined as the amount of enzyme which releases 1 μmol of xylose in 1 min under assay conditions.
Cellulase assay
Cellulase activity (CMCase and FPase) was analysed as described by Wood and Bhat (1988). CMCase activity was determined by measuring the reducing sugars produced from 1% (w/v) carboxymethylcellulose, whereas FPase activity was determined by measuring the reducing sugar from Whatman filter paper No. 1. The liberated reducing sugars were measured using the dinitrosalicylic acid (DNS) method (Miller 1959). One unit of CMCase and FPase activity is defined as the amount of enzyme which releases 1 μmol of glucose in 1 min under assay conditions.
Protein assay
The protein content of the sample was determined according to the Bradford method (Bradford 1976), using bovine serum albumin as the standard (Sigma-Aldrich Catalog).
RESULTS AND DISCUSSION
Screening for Fungal Xylanase Producer
Among the four fungi tested, P. oxalicum T3.3 and A. niger ATCC 6275 produced the largest clear zones based on the relative enzyme activity determination. In contrast, C. gloeosporioides and P. sanguineus did not produce any clear zones. From another study, Gutierrez-Soto et al. (2015) revealed that Pycnoporus sanguineus can produce a clear zone by using plate screening test, in contrast with this study. The fungus showed the highest xylanase activity, corresponding to 18.39 U/mL, which was obtained in submerged fermentation using a combination of wheat straw and orange peel as the substrate. Table 1 shows the relative enzyme activity for each species based on the plate screening test. The highest relative enzyme activity was obtained from P. oxalicum T3.3 (2.01 ± 0.39), followed by A. niger ATCC 6275 (1.2 ± 0.13). This result is in agreement with results reported by Polizeli et al. (2005), where Aspergillus and Penicillium sp. have the most commercial potential as xylanase producers among the filamentous fungi. Currently, the qualitative screening that uses a solid screening medium containing xylan as the sole substrate is popular among researchers to identify xylanase producers based on the diffusion principle. This halo detection is an easy and valuable way of conducting screening (Ribeiro et al. 2014), where the Congo red (used for staining) has specific link and binds strongly with xylan containing β-1,4-xylosidic linkages (Samanta et al. 2011). The degradation of these linkages rendered the Congo red unable to associate with xylan or to produce a clear zone on the medium after repeatedly cleansing with 1 M NaCl. Although this method saves times and energy, it also has limitations. This technique is dependent on the fungus, where some fungi may sporulate too much and spread over the plates, making it difficult to interpret the results (Ribeiro et al. 2014). Therefore, the growth of fungus needs to be monitored daily to ensure that the fungus will not spread over the plate. Hence, the diameter of the mycelial and clear zone can be determined. The diameter of clear zone varies depending on the fungal species.
The result from the clear zone analysis only gave a qualitative result of the hydrolytic enzyme production and is not an accurate method to determine the maximum enzyme production. This is because there are some technical problems that can influence the clear zone size. For example, in addition to the growth kinetics of the isolates, the migration of enzyme released depends upon the agar percentage, molecular size and weight of the enzyme, and growth temperature (Sridevi and Charya 2011). Therefore, the quantitative screening by submerged fermentation needs to be conducted to distinguish the most efficient xylanase enzyme-producers among the fungi species that produce clear zones.
Table 1. Relative Enzyme Activities of Fungi Species on the Screening Plate Based on the Clear Zone Formation
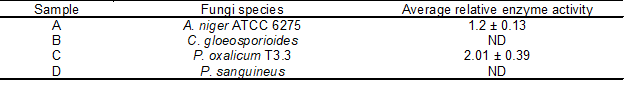
ND: not detected
Time Course of Xylanase and Cellulase Enzyme Production by Submerged Fermentation
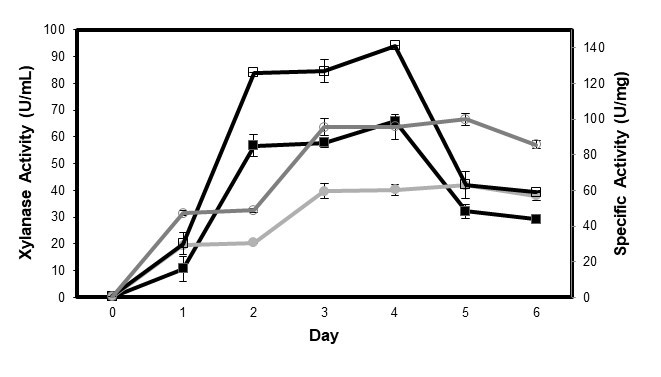
The positive fungi species (A. niger ATCC 6275 and P. oxalicum T3.3) were selected for this stage of study. The two species were used to determine xylanase production via submerged fermentation after six days of incubation. The results in Fig. 1 show the amount of reducing sugars released by the enzyme of P. oxalicum T3.3 and A. niger ATCC 6275. The xylanase activities started on the first day of incubation for both species of fungi. For P. oxalicum T3.3, maximum xylanase production was observed after four days of incubation with an enzyme activity of 65.89 U/mL and 141.14 U/mg proteins. The enzyme activity pattern obtained from P. oxalicum T3.3 shows slight increase in enzyme production from day 0 to the first day. The pattern changed and gradually increased from the second day to the fourth day. A noticeable decrease in xylanase activities was seen from the fifth day onwards. Meanwhile, maximum xylanase production by A. niger ATCC 6275 occurred on the fifth day of fermentation, with an enzyme activity of 41.95 U/mL and 99.84 U/mg proteins. The result showed that P. oxalicum T3.3 needed less incubation time and produced higher enzyme activity compared to A. niger ATCC 6275. Earlier reports indicated the maximum xylanase production by fungi occurred after 4 to 5 days of fermentation (Carmona et al. 1998; Ruckmani and Rajendran 2001; Shah and Madamwar 2005; Murthy and Naidu 2010). The results of this study are in good agreement with the results obtained by Bajaj et al. (2011), where the highest xylanase production from Penicillium sp. SS1 was reported on the fourth day (43.84 U/mL). Furthermore, the highest xylanase production for A. terreus was also captured on the fourth day (Sathish et al. 2008), and A. niger KKS was captured on the sixth day of fermentation, using rice straw as the substrate (Kang et al. 1995). According to Bajaj et al. (2011), the optimum fermentation period of any organism is dependent on the microbial strain and its genetic make-up as well as on cultural and environmental conditions during the growth of the organism. The decrease in enzyme activity was due to the release of xylanase from the bound substrate and from the autolysis of fungal cells (Bushan et al. 2012). Therefore, P. oxalicum T3.3 offers a better choice compared to A. niger ATCC 6275. This is because P. oxalicum T3.3 reduces the risk of contamination and the cost of enzyme production as a result of its shorter incubation time.
Fig. 1. Time course of xylanase enzyme production and specific activity by P. oxalicum T3.3 and A. niger ATCC 6275. Values were derived after three replicates with ± SD. Symbol represent :![]() xylanase activity from P. oxalicum T3.3;
xylanase activity from P. oxalicum T3.3; ![]() xylanase activity from A. niger ATCC 6275;
xylanase activity from A. niger ATCC 6275; ![]() specific activity from P. oxalicum T3.3;
specific activity from P. oxalicum T3.3; ![]() specific activity from A. niger ATCC 6275
specific activity from A. niger ATCC 6275
Because of these advantages, P. oxalicum T3.3 was chosen for the next stage of study. Although the high xylanase enzyme was important in the biobleaching process, the level of cellulase enzyme also needs to be considered to avoid the negative effects. The cellulase activity for CMCase and FPase enzyme production from P. oxalicum T3.3, using rice straw as a substrate, was determined. In many applications, the xylanase enzyme must be free of cellulase enzyme activity or have very low traces of it (Nagar et al. 2013). In fact, in biobleaching application, cellulase activity gives many negative effects such as loss of cellulose, degradation of pulp quality, and increased effluent treatment cost (Beg et al. 2001). Figure 2 shows that P. oxalicum T3.3 had very low cellulase enzyme activity, indicating that the xylanase was more or less cellulase-free.
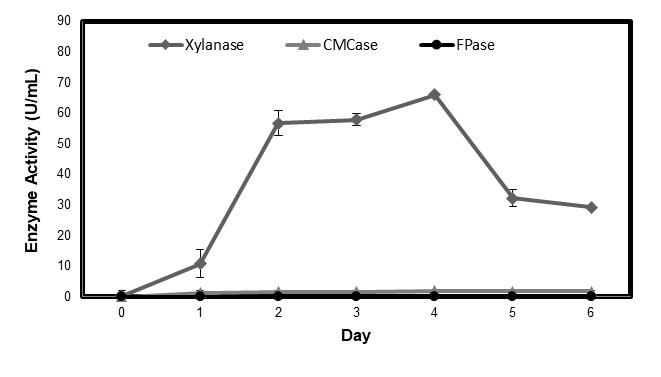
Fig. 2. Production of xylanase, CMCase, and FPase enzyme from P. oxalicum T3.3. Values are means of 3 replicates with ± SD.
Table 2. Enzyme Yields under Submerged Fermentation by Various Isolates and Carbon Sources
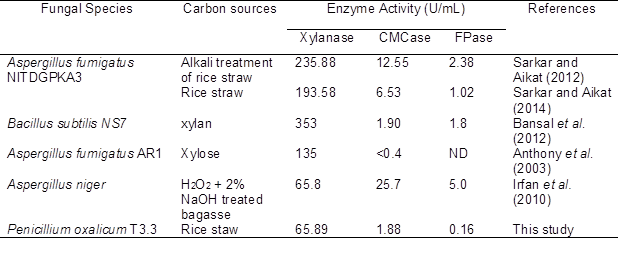
ND: not detected
The highest production for both enzymes was captured on the fourth day of fermentation, 1.88 U/mL for CMCase and 0.16 U/mL for FPase. This shows that rice straw induced the production of xylanase by P. oxalicum T3.3 in comparison to cellulase production. The comparison of xylanase, CMCase, and FPase enzymes production from this study, as well as findings by other references, are presented in Table 2.
Rice Straw vs. Beechwood Xylan as Substrate
Figure 3 shows xylanase production from P. oxalicum T3.3 using rice straw and beechwood xylan as substrates. The production increased after four days of fermentation, but it dropped gradually on the 5th and 6th day. The xylanase production was fairly similar for these substrates: 65.89 U/mL (141.14 U/mg protein) for rice straw and 64.72 U/mL (119.81 U/mg protein) for beechwood xylan. Although beechwood xylan can support xylanase production, the results from this experiment indicate that the medium containing rice straw yielded higher xylanase enzymes compared to the medium containing beechwood xylan at earlier periods of fermentation. Because the production of xylanase enzymes is affected by the substrate, the choice of an appropriate substrate is of great importance in having successful xylanase production. This is because the substrate not only serves as a carbon source, but also as an inducer for enzyme production (Bakri et al. 2010). Therefore, it can be concluded that P. oxalicum T3.3 performed better in media containing rice straw as the substrate for the growth and production of enzymes. In any industrial production process, the cost of the substrate plays a crucial role. Therefore, the low-value lignocellulosic waste was preferred compared to an expensive substrate (commercial xylan) for large-scale xylanase production (Bakri et al. 2010; Bajaj et al. 2011). Besides, this study was done in Malaysia where rice is known as a major food for Malaysians. That means, rice straw are abundance in Malaysia and these rice straw can be used as a valuable energy source for enzyme production. Additionally, utilization of lignocellulosic wastes also helps to solve the environmental waste disposal problem (Bajaj et al. 2011). Besides the cost of production, we also can protect the environment from the pollution with using this lignocellulosic waste.
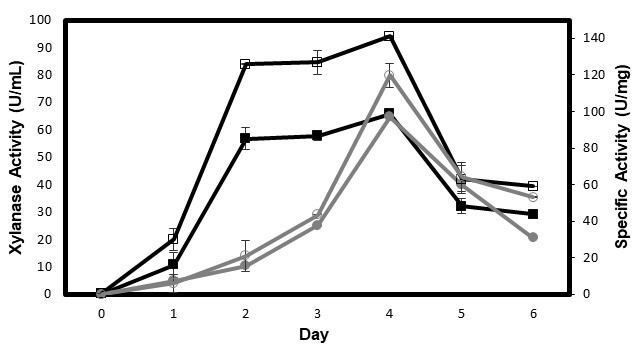
Fig. 3. Effects of rice straw and beechwood xylan on xylanase enzyme production. Values are means of 3 replicates with ± SD. Symbol represent:![]() xylanase activity using rice straw;
xylanase activity using rice straw;![]() xylanase activity using beechwood xylan;
xylanase activity using beechwood xylan; ![]() specific activity using rice straw;
specific activity using rice straw; ![]() specific activity using beechwood xylan
specific activity using beechwood xylan
Table 3 shows a comparison of xylanase production by various species of fungi using various agro-residues and commercial xylans. The production of xylanase by P. oxalicum T3.3 in this study was comparable with other findings.
Table 3. Xylanase Production under Submerged Fermentation using Different Lignocellulosic Waste and Commercial Xylan as Substrate
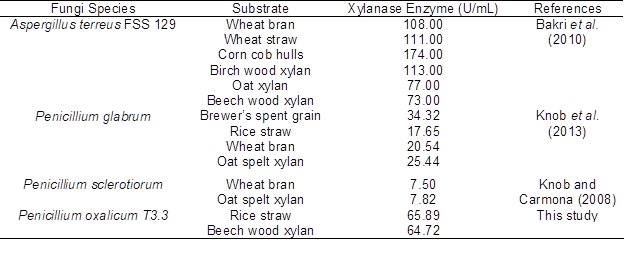
The differences in chemical composition of lignocellulosic wastes and the accessibility of the substrates to the fungi are important factors that affect the production level of enzyme (Knob et al. 2013). Therefore, rice straw can be seen as an acceptable substrate for producing xylanase using P. oxalicum T3.3.
CONCLUSIONS
- oxalicum T3.3 was found to be a potential xylanase producer with low cellulase enzyme activity.
- Rice straw is a suitable substrate for the production of xylanase.
ACKNOWLEDGMENTS
The authors would like to acknowledge the Universiti Putra Grant Scheme for (Project No.: GP-IPB/2013/9413/9413402), and a Graduate Research Fellowship (GRF) from Universiti Putra Malaysia for supporting this research.
REFERENCES CITED
Abdel-Sater, M. A., and El-Said, A. H. M. (2001). “Xylan-decomposing fungi and xylanolytic activity in agricultural and industrial wastes,” International Biodeterioration and Biodegradation 47(1), 15-21. DOI: 10.1016/S0964-8305(00)00113-X
Abu, E. A., Onyenekekwe, P. C., Ameh, D. A., Agbaji, A. S., and Ado, S. A. (2000). “Cellulase (E.C.3.2.1.3) production from sorghum bran by Aspergillus niger SL1: An assessment of pretreatment method,” Proceedings of the International Conference on Biotechnology: Commercialization and Food security, Abuja, Nigeria, pp. 153-159.
Adesina, F. C., and Onilude, A. A. (2013). “Isolation, identification and screening of xylanase and glucanase-producing microfungi from degrading wood in Nigeria,” African Journal of Agricultural Research 8(34), 4414-4421.
Ali, M., and Sreekrishnan, T. R. (2001). “Aquatic toxicity from pulp and paper mill effluents: a review,” Advances in Environmental Research 5(2), 175-196. DOI: 10.1016/S1093-0191(00)00055-1
Anthony, T., Raj, K. C., Rajendran, A., and Gunakesaran, P. (2003). “High molecular weight cellulase-free xylanase from alkali-tolerant Aspergillus fumigatus AR1,” Journal of Bioscience and Bioengineering 96(4), 394-396. DOI: 10.1016/S1389-1723(03)90143-5
Bajaj, B. K., Sharma, M., and Sharma, S. (2011). “Alkalistable endo-β-1,4-xylanase production from a newly isolated alkalitolerant Penicillium sp. SS1 using agro-residues,” 3 Biotech 1(2), 83-90. DOI: 10.1007/s13205-011-0009-5
Bakri, B., Al-Jazairi, M., and Al-Kayat, G. (2008). “Xylanase production by a newly Aspergillus niger SS7 in submerged culture,” Polish Journal of Microbiology 57(3), 249-251.
Bakri, Y., Masson, M., and Thonart, P. (2010). “Isolation and identification of two new fungal strains for xylanase production,” Applied of Biochemistry and Biotechnology 162(6), 1626-1634. DOI: 10.1007/s12010-010-8944-x
Bansal, N., Raman, S., Janvega, C., and Soni, S. K. (2012). “Production of xylanase-cellulase complex Bacillus subtilis NS7 for the biodegradation of agrowaste residues,” Lignocellulose 1(3), 196-209.
Beg, Q. K., Bhushan, B., Kapoor, M., and Hoondal, G. S. (2000). “Enhanced production of a thermostable xylanase from Streptomyces sp. QG-11-3 and its application in biobleaching of eucalyptus kraft pulp,” Enzyme and Microbial Technology 27(7), 459-466. DOI: 10.1016/S0141-0229(00)00231-3
Beg, Q. K., Kapoor, M., Mahajan, L., and Hoondal, G. S. (2001). “Microbial xylanases and their industrial applications: A review,” Applied Microbiology and Biotechnology 56(3-4), 326-338. DOI: 10.1007/s002530100704
Bradford, M. M. (1976). “A rapid and sensitive method for quantitation of microgram quantities of protein utilizing the principle of protein-dye binding,” Analytical Biochemistry 72 (1-2), 248-254. DOI: 10.1016/0003-2697(76)90527-3
Bushan, B., Pal, A., and Jain, V. (2012). “Isolation, screening and optimized production of extracellular xylanase under submerged condition from Aspergillus flavus Mtcc 9390,” Enzyme Engineering 1(2), 1-6. DOI: 10.4172/eeg.1000103
Carmona, E. C., Brochetto-braga, M. R., Pizzirani-kleiner, A. A., and Jorge, J. A. (1998). “Purification and biochemical characterization of an endoxylanase from Aspergillus versicolor,” FEMS Microbiology Letters 166(2), 311-315. DOI: 10.1111/j.1574-6968.1998.tb13906.x
Collins, T., Gerday, C., and Feller, G. (2005). “Xylanases, xylanase families and extremophilic xylanases,” FEMS Microbiology Reviews 29(1), 3-23. DOI: 10.1016/j.femsre.2004.06.005
De Ioannes, P., Peirano, A., Steiner, J., and Eyzaguirre, J. (2000). “An α-L-arabinofuranosidase from Penicillium purpurogenum: Production, purification and properties,” Journal of Biotechnology 76(2-3), 253-258. DOI: 10.1016/S0168-1656(99)00190-X
Egaña, L., Gutiérrez, R., Caputo, V., Peirano, A., Steiner, J., and Eyzaguirre, J. (1996). “Purification and characterization of two acetyl xylan esterases from Penicillium purpurogenum,” Biotechnology and Applied Biochemistry 24(1), 33-39.
Eyzaguirre, J., Scarpa, J., Belancic, A. and Steiner, J. (1992). “The xylanase system of Penicillium purpurogenum,” in: Xylans and Xylanases, Visser, J. G., Beldman, M. A., van Someren, K., and Voragen, A. G. J. (eds.), Elsevier, Amsterdam, pp. 505-510.
Food and Agriculture Organization (FAO) (2010). Available from: http://faostat.fao.org. (accessed October 1, 2014).
Food and Agriculture Organization (FAO) (2014). Available from: http://faostat.fao.org. (accessed October 1, 2014).
Ghatora, S. K., Chadia, B. S., Bhadan, A. K., and Bhat, M. K. (2006). “Identification and characterisation of diverse xylanases from thermophilic and thermotolerant fungi,” BioResources 1(1), 18-33. DOI: 10.15376/biores.1.1.18-33
Gupta, S., Bhushan, B., and Hoondal, G. S. (2000). “Isolation, purification and characterization of xylanase from Staphylococcus sp. SG-13 and its application in biobleaching of kraft pulp,” Journal of Applied Microbiology 88(2), 325-334. DOI: 10.1046/j.1365-2672.2000.00974.x
Gutierrez-Soto, G., Medina-Gonzalez, G. E., Garcia-Zambrano, E. A., Trevino-Ramirez, J. E., and Hernendez-Luna, C. E. (2015). “Selection and characterization of a native Pycnoporus sanguineus strain as a lignocellulolytic extract producer from submerged cultures of various agroindustrial wastes,” BioResources 10(2), 3564-3576.
Harris, A.D. and Ramalingam, C. (2010). “Xylanases and its application in food industry: A review,” Journal of Experimental Sciences 1(7), 1-11.
Hatanaka, K. (2012). “Incorporation of fluorous glycosides to cell membrane and saccharide chain elongation by cellular enzymes,” Topics in Current Chemistry 308(Special Issue), 291-306. DOI: 10.1007/128_2011_276
Henriksson, G., and Teeri, T. (2009). “Biotechnology in the forest industry,” In: Pulp and Paper Chemistry and Technology Volume 1. Wood Chemistry and Wood Biotechnology, Ek, M., Gellerstedt, G., and Henrikksson, G. (eds.), Walter de Gruyter, pp. 273-300.
Howard, R. L., Abotsi, E., Jansen van Rensburg, E. L., and Howard, S. (2003). “Lignocellulose biotechnology: Issues of bioconversion and enzyme production, Review,” African Journal of Biotechnology 2(12), 602-619. DOI: 10.5897/AJB2003.000-1115
Irfan, M., Nadeem, M., Syed, Q. A., and Baig, S. (2010). “Submerged cultivation of Aspergillus niger on pretreated sugarcane bagasse,” World Journal of Agricultural Sciences 6(4), 466-472.
Kang, S. W., Kim, S. W., and Lee, J. S. (1995). “Production of cellulase and xylanase in a bubble column using immobilized Aspergillus niger KKS,” Applied Biochemistry and Biotechnology 53(2), 101-106. DOI: 10.1007/BF02788601
Kanimozhi, K., and Nagalakshmi, P. K. (2014). “Xylanase production from Aspergillus niger by solid state fermentation using agricultural waste as substrate,” International Journal of Current Microbiology and Applied Sciences 3(3), 437-446.
Knob, A., and Carmona, E. C. (2008). “Xylanase production by Penicillium sclerotiorum and its characterization,” World Applied Sciences Journal 4(2), 277-283.
Knob, A., Beiltel, S. M., Fortkamp, D., Terrasan, C. R. F. and Almelda, A. F. D. (2013). “Production, purification, and characterization of a major Penicillium glabrum xylanase using brewer’s spent grain as substrate,” BioMed Research International 2013, 728735. DOI: 10.1155/2013/728735
Koponen, R. (1991). “Enzyme systems prove their potential,” Pulp and Paper International 33(11), 81-83.
Kulkarni, N., Shendye, A., and Rao, M. (1999). “Molecular and biotechnological aspects of xylanases,” FEMS Microbiology Reviews 23(4), 411-456. DOI: 10.1111/j.1574-6976.1999.tb00407.x
Latif, F., Asgher, M., Saleem, R., Akrem, A., and Legge, R. L. (2006). “Purification and characterization of a xylanase produce by Chaetomium thermophile NIBGE,” World Journal of Microbiology and Biotechnology 22(1), 45-50. DOI: 10.1007/s11274-005-5745-4
Lembaga Kemajuan Pertanian Muda (MADA) (2010) Laporan Tahunan, Alor Setar, Kedah.
Levasseur, A., Asther, M., and Record, E. (2005). “Overproduction and characterization of xylanase B from Aspergillus niger,” Canadian Journal of Microbiology 51(2), 177-183. DOI: 10.1139/w04-116. DOI: 10.1139/w04-116
Li, K., Azadi, P., Collins, R., Tolan, J., Kim, J. S., and Eriksson, K.-E. L. (2000) “Relationships between activities of xylanases and xylan structures,” Enzyme and Microbial Technology 27(1-2), 89-94. DOI: 10.1016/S0141-0229(00)00190-3
Liao, H., Zheng, H., Li, S., Wei, Z., Mei, X., Ma, H., Shen, Q. and Xu, Y. (2015). “Functional diversity and properties of multiple xylanases from Penicillium oxalicum GZ2,” Scientific Reports 5:12631. DOI: 10.1038/srep12631
Mandels, M., Hontz, I., and Nystrom, J. (1974). “Enzymatic hydrolysis of waste cellulose,” Biotechnology and Bioengineering 16(11), 1471-1493. DOI: 10.1002/b.it.260161105
Miller, G., L. (1959). “Use of dinitrosalicylic acid reagent for determination of reducing sugar,” Analytical Chemistry 31(3), 426-428.
McCrae, I. S., Leith, K. M., Gorden, A. H., and Wood, T. M. (1994). “Xylan-degrading enzyme system produced by fungus Aspergillus awamori, isolation and characterization of a feruloyl esterase and a pcoumaroyl esterase,” Enzyme and Microbial Technology 16(10), 826-834. DOI: 10.1016/0141-0229(94)90055-8
Murthy, P. S., and Naidu, M. M. (2010). “Production and application of xylanase from Penicillium sp. utilizing coffee by-products,” Food and Bioprocess Technology 5(2), 657-664. DOI: 10.1007/s11947-010-0331-7
Nagar, S., Jain, R. K., Thakur, V. V., and Gupta, V. J. (2013). “Biobleaching application of cellulose poor and alkaline stable xylanase from Bacillus pumilus SV-85S,” 3 Biotech. 3(4), 277-285. DOI: 10.1007/s13205-012-0096-y
Nakamura, S. (2003). “Structure and function of a multidomain alkaline xylanase from alkaliphilic Bacillus sp. strain 41M-1,” Catalysis Surveys from Asia 7(2-7), 157-164. DOI: 10.1023/A:1025337608799
Norazlina, I., Meenalosani, N., and KuHalim, K. H. (2013). “Production of xylanase by Trichoderma sp. via solid state culture using sugarcane bagasse,” International Journal of Energy Science 3(2), 99-105.
Okafor, U. A., Okochi, V. I., Onyegeme-okereta, B. M., and Nwodo-Chinedu, S. (2007). “Xylanase production by Aspergillus niger ANL 301 using agro-wastes,” African Journal of Biotechnology 6(14), 1710-1714.
Paloheimo, M., Mantyla, A., and Kallio, K. (2003). “High-yield production of a bacterial xylanase in the filamentous fungus Trichoderma reesei requires a carrier polypeptide with an intact domain structure,” Applied and Environmental Microbiology 69(12), 7073-7082. DOI: 10.1128/AEM.69.12.7073-7082.2003.
Pang, P. K., and Ibrahim, C. O. (2005). “Xylanase production by a local fungal isolate, Aspergillus niger USM AI 1 via solid state fermentation using palm kernel cake (PKC) as substrate,” Songklanakar Journal of Science and Technology 27(2), 325-336.
Polizeli, M. L. T. M., Rizzatti, A. C. S., Monti, R., Terenzi, H. F., Jorge, J. A., and Amorim, D. S. (2005). “Xylanases from fungi: Properties and industrial applications,” Applied Microbiology and Biotechnology 67(5), 577-591. DOI: 10.1007/s00253-005-1904-7
Ribeiro, L. F. C., Ribeiro, L. F, Jorge, J. A., and Polizeli, M. L. T. M. (2014). “Screening of filamentous fungi for xylanases and cellulases not inhibited by xylose and glucose,” British Biotechnology Journal 4(1), 30-39. DOI: 10.9734/BBJ/2014/6066
Ruckmani, A., and Rajendran, A. (2001). “Production of cellulase-poor xylanase by an alkalitolerant strain of Aspergillus flavus,” Indian Journal of Microbiology 41, 115-118.
Sahin, H. L., and Mantanis, G. I. (2011). “Colour changes in wood surfaces modified by a nanoparticulate based treatment,” Wood Research 56(4), 525-532
Samanta, A. K., Kolte, A., Senani, S., Sridhar, M., and Jayapal, N. (2011). “A simple and efficient diffusion technique for assay of endo β-1, 4-xylanase activity,” Brazilian Journal of Microbiology 42(4), 1349-1353. DOI: 10.1590/S1517-83822011000400016
Sarkar, N., and Aikat, K. (2012). “Alkali pretreatment of rice straw and enhanced cellulase production by a locally isolated fungus Aspergillus fumigatus NITDGPKA3,” Journal of Microbiology and Biotechnology Research 2(5), 717-726.
Sarkar, N., and Aikat, K. (2014). “Aspergillus fumigatus NITDGKA3 provides for increased cellulase production,” International Journal of Chemical Engineering 2014, 959845. DOI: 10.1155/2014/959845
Sathish, T., Lakshmi, G. S., Roa, C. H. S., Brahmaiah, P., and Prakasham, R. S. (2008). “Mixture design as first step for improved glutaminase production in solid-state fermentation by isolated Bacillus,” Letters in Applied Microbiology 47(4), 256-262. DOI: 10.1111/j.1472-765X.2008.02413.x
Shah, A. R., and Madamwar, D. (2005). “Xylanase production by a newly isolated A. foetidus strain and its characterization,” Process Biochemistry 40(5), 763-777. DOI: 10.1016/j.procbio.2004.06.041
Sridevi, B., and Charya, M. A. S. (2011). “Isolation, identification and screening of potential cellulose-free xylanase producing fungi,” African Journal of Biotechnology 10(22), 4624-4630.
Subramaniyan, S., and Prema, P. (2002). “Biotechnology of microbial xylanases: Enzymology, molecular biology and application,” Critical Reviews in Biotechnology 22(1), 33-64. DOI: 10.1080/07388550290789450
Sun, W. C., Cheng, C. H., and Lee, W. C. (2008). “Protein expression and enzymatic activity of cellulases produced by Trichoderma reesei Rut C-30 on rice straw,” Process Biochemistry 43(10), 1083-1087. DOI: 10.1016/j.procbio.2008.05.015
Techapun, C., Poosaran, N., Watanabe, M., and Sasaki, K. (2003). “Thermostable and alkaline-tolerant microbial cellulase-free xylanases produced from agricultural wastes and the properties required for use in pulp bleaching bioprocesses: A review,” Process Biochemistry 38(9), 1327-1340. DOI: 10.1016/S0032-9592(02)00331-X
United States Department of Agriculture (USDA) (2014). Available from: http://worldriceproduction.com (accessed June 24, 2015).
Wong, K. K. Y., Tan, L. U. L., and Saddler, J. N. (1988). “Multiplicity of B-1,4-xylanase in microorganisms: Functions and applications,” Microbiological Reviews 52(3), 305-317.
Wood, T. M., and Bhat, K. M. (1988). “Methods for measuring cellulase activities,” Methods in Enzymology 160, 87-112
Article submitted: May 20, 2015; Peer review completed: September 5, 2015; Revised version received and accepted: November 17, 2015; Published: December 10, 2015.
DOI: 10.15376/biores.11.1.1162-1175
